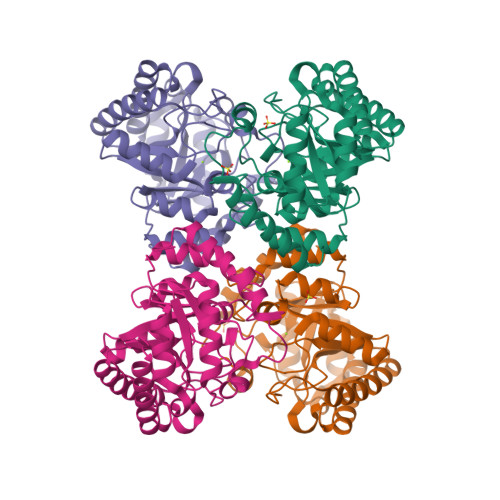
3D Structure View
Loading...
More Info
GENERAL
TITLE
CRYSTAL STRUCTURES OF D-ALLULOSE 3-EPIMERASE FROM SINORHIZOBIUM FREDII
DATE LAST MODIFIED
-
RELEASE
3 Aug 2022
OBSOLETE
CLASSIFICATION
ISOMERASE
EXPERIMENT TYPE
X-RAY DIFFRACTION
SPACE GROUP
P 1 21 1
RESOLUTION RANGE HIGH
1.55
RESOLUTION RANGE LOW
45.17
PUBMED ID
-
DOI
--
PDBe
--
LIGANDS
--
Chains
| Structure Chain | Target Protein | Gene | Expression System | Organism | Synonims | Actions |
|---|---|---|---|---|---|---|
| Q16186 | SF83666_B55120 | ESCHERICHIA COLI BL21(DE3) | SINORHIZOBIUM FREDII CCBAU 83666 | -- | ||
| P0CG47 | SF83666_B55120 | ESCHERICHIA COLI BL21(DE3) | SINORHIZOBIUM FREDII CCBAU 83666 | -- | ||
| Q99460 | SF83666_B55120 | ESCHERICHIA COLI BL21(DE3) | SINORHIZOBIUM FREDII CCBAU 83666 | -- | ||
| Q99460 | SF83666_B55120 | ESCHERICHIA COLI BL21(DE3) | SINORHIZOBIUM FREDII CCBAU 83666 | -- |
NGL Viewer is used for the molecular visualization.
- AS Rose, AR Bradley, Y Valasatava, JM Duarte, A Prlić and PW Rose. NGL viewer: web-based molecular graphics for large complexes. Bioinformatics: bty419, 2018. doi:10.1093/bioinformatics/bty419
- AS Rose and PW Hildebrand. NGL Viewer: a web application for molecular visualization. Nucl Acids Res (1 July 2015) 43 (W1): W576-W579 first published online April 29, 2015. doi:10.1093/nar/gkv402