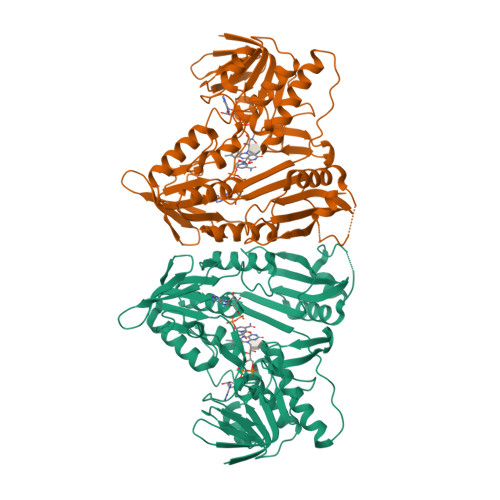
3D Structure View
Loading...
More Info
GENERAL
TITLE
STRUCTURE AND CATALYTIC MECHANISM OF MONODEHYDROASCORBATE REDUCTASE, MDHAR, FROM ORYZA SATIVA L. JAPONICA
DATE LAST MODIFIED
29 Jul 2020
RELEASE
12 Oct 2016
OBSOLETE
CLASSIFICATION
OXIDOREDUCTASE
EXPERIMENT TYPE
X-RAY DIFFRACTION
SPACE GROUP
P 21 21 21
RESOLUTION RANGE HIGH
2.29
RESOLUTION RANGE LOW
71.76
PUBMED ID
DOI
10.1038/SREP33903
PDBe
--
Chains
| Structure Chain | Target Protein | Gene | Expression System | Organism | Synonims | Actions |
|---|---|---|---|---|---|---|
| B6YWC1 | OS09G0567300, OS09G0567300, OJ1155_H10.27, OSNPB_090567300 | ESCHERICHIA COLI | RICE | PUTATIVE MONODEHYDROASCORBATE REDUCTASE | ||
| Q652L6 | OS09G0567300, OS09G0567300, OJ1155_H10.27, OSNPB_090567300 | ESCHERICHIA COLI | RICE | PUTATIVE MONODEHYDROASCORBATE REDUCTASE | Mutations |
NGL Viewer is used for the molecular visualization.
- AS Rose, AR Bradley, Y Valasatava, JM Duarte, A Prlić and PW Rose. NGL viewer: web-based molecular graphics for large complexes. Bioinformatics: bty419, 2018. doi:10.1093/bioinformatics/bty419
- AS Rose and PW Hildebrand. NGL Viewer: a web application for molecular visualization. Nucl Acids Res (1 July 2015) 43 (W1): W576-W579 first published online April 29, 2015. doi:10.1093/nar/gkv402